...
- Install the latest version of R from http://cran.r-project.org/bin/windows/base/ with default settings. Under the default settings, R should be installed in the "C:\Program Files\R" directory.
Run R in interactive mode by executing R.exe available at "C:\Program Files\R\R-3.1.1\bin\R.exe" and install the various necessary packages by entering the following commands in the command prompt:
install.packages("ggplot2")install.packages("gstat")
install.packages("moments")
install.packages("fields")
install.packages("GA")
install.packages("spdep")
install.packages("rgeos")
install.packages("fields")
- Download the code from https://uofi.box.com/s/g354lm5av36uz0v27q75 Extract the zipped folder and you should be able to see the contents as shown below.
Execution - Execute the application by double clicking the 'Triaxus Script' shortcut.
- Select the main directory for the application where all the files exist as the working directory, by clicking on the Set working directory button.
- Choose the data file to be used by clicking on the browse button to choose an appropriate source data file.
- Select the various Physical and/or Biological features that you want the script to consider, by checking the appropriate boxes adjacent to each of the features.
- Click the 'Run script' button to start the script execution.
- The Console window would give real time status updates on the script's execution.
- Sample Outputs generated from the Triaxus Script for the Temperature and SUNA Nitrate features Triaxus Script Output : Feature = Temperature , Triaxus Script : Feature = SUNA Nitrate
Instructions for running the Seabird script on a windows machine
Steps
- Install Python version 2.7 from https://www.python.org/download/releases/2.7.7/
Install the scikit-learn machine learning library available as "scikit‑learn‑0.15.1.win32‑py2.7.exe" at
http://www.lfd.uci.edu/~gohlke/pythonlibs/#scikit-learnInstalling the Scipy stack for python 2.7 available as "Scipy-stack-14.5.30.win32-py2.7.exe" at
http://www.lfd.uci.edu/~gohlke/pythonlibs/#scipy-stack- Download the code from https://uofi.box.com/s/h6pvnackfmvq3kstzjzn Extract the zipped folder and you should be able to see the contents as shown below.
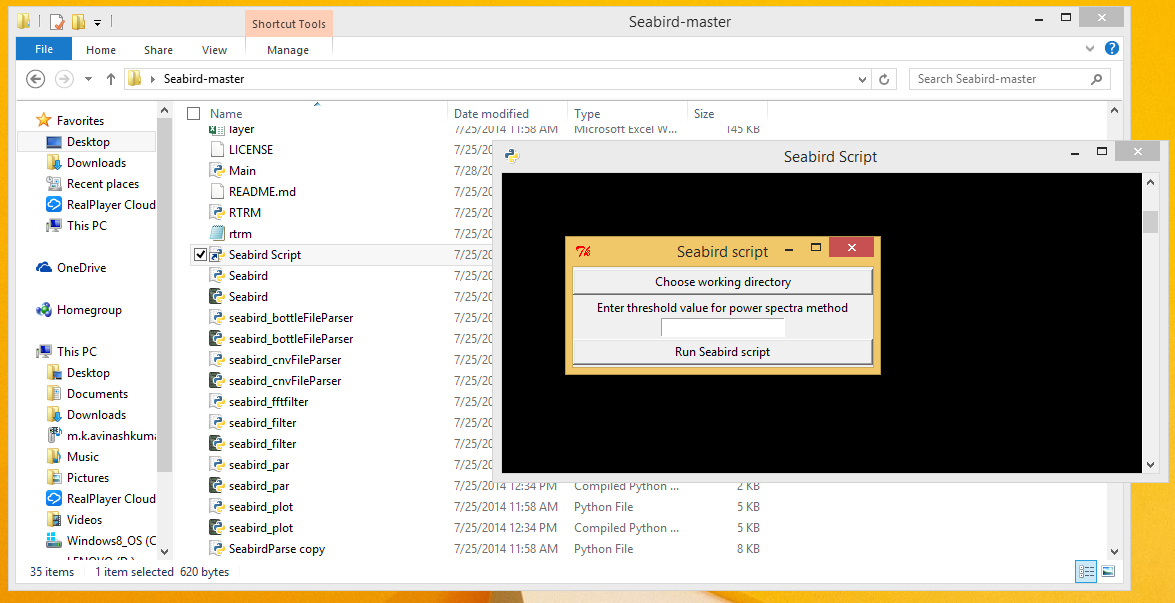
- Execute the application by double clicking on the 'Seabird Script' shortcut.
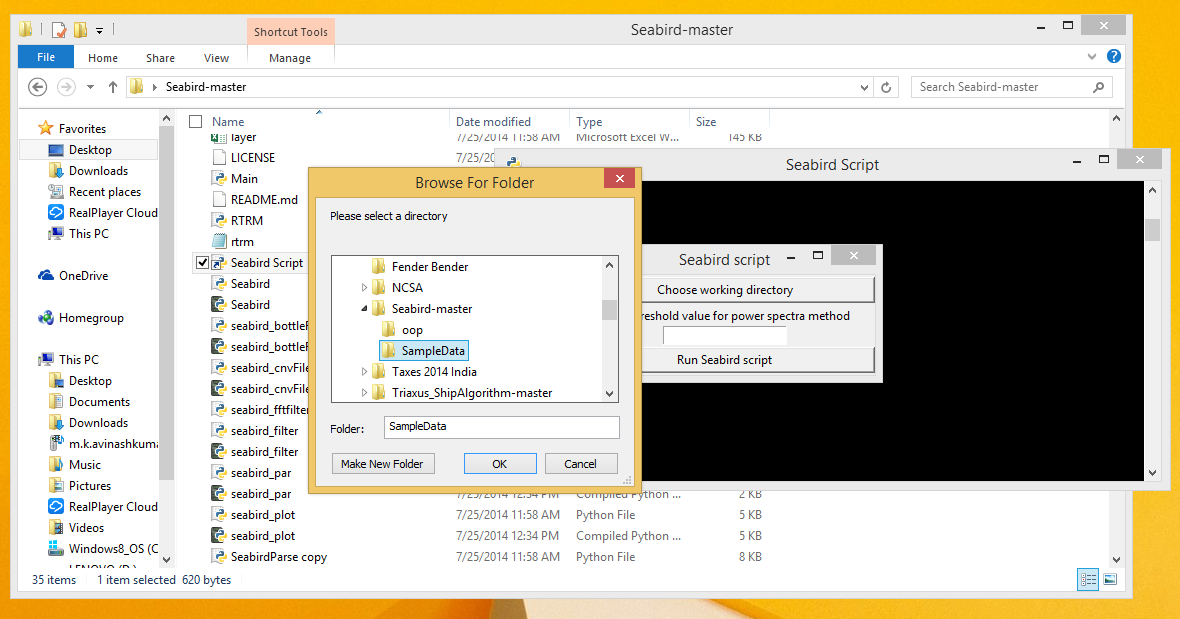
- Choose the directory where all the input data files as the working directory by clicking on the 'Choose working directory' button.
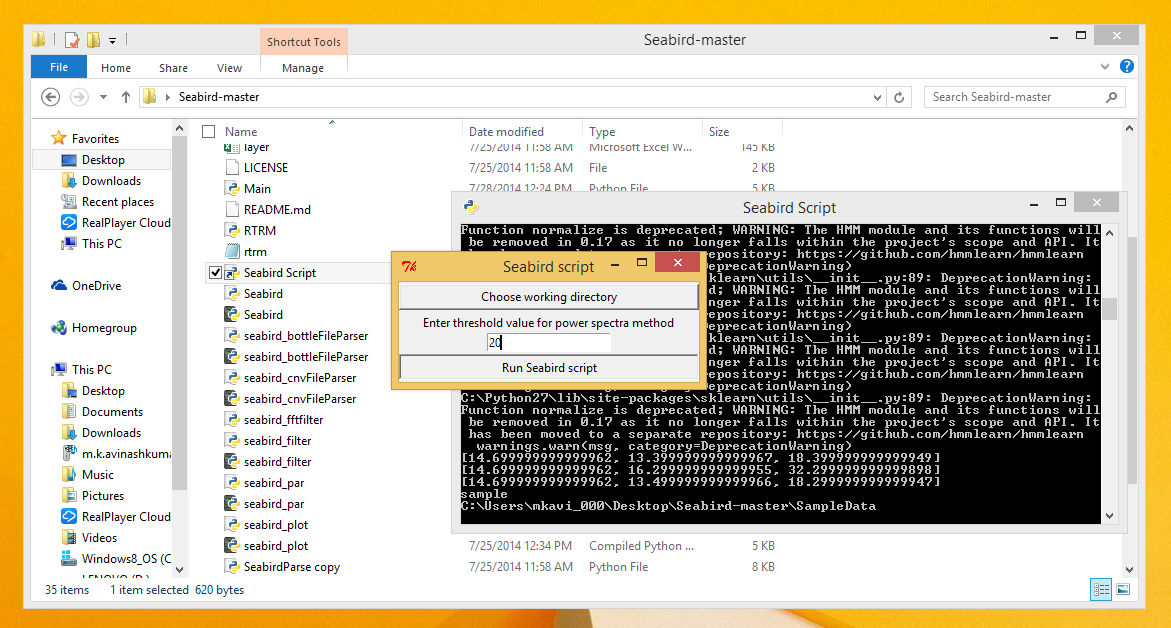
- Enter a threshold value for the Power Spectra method if necessary and Click the 'Run Seabird script' button.
- Once the Script finishes execution, the output files should have been generated in the folder designated as the working directory.
...
- Install R
- sudo apt-get install r-base
- sudo apt-get install libgeos++-dev
- Go to the R execution environment by entering sudo R
- Install various prerequisite packages through the following commands
install.packages("ggplot2")
install.packages("gstat")
install.packages("moments")
install.packages("fields")
install.packages("GA")
install.packages("spdep")
- Create a Folder called Result inside the applications directory and a sub folder called Variogram inside the newly created Result folder. These folders would be used by the script to store some meta data.
- Run the main entry R script for the application, by entering source("Main.R")
- The final output graph should be available as a pop up on the terminal.
Instructions for running the
...
Triaxus R script on a linux machine
Download the seabird code from https://github.com/stormxuwz/Seabird
...